Introducing BAMM: A Standardized Metadata Model for Clinical Chart Abstraction
March 16, 2026
In healthcare, chart abstraction quietly powers some of the most consequential decisions we make for patients. Although it rarely gets the spotlight, abstraction is the connective tissue across research, quality measurement, and clinical operations as the process by which critical facts buried in clinical notes become structured, actionable data.
And today, as AI reshapes how organizations extract value from those notes, the need for a standardized framework for defining and validating abstraction has never been more urgent.
At Brim, we believe abstraction should be transparent, reproducible, interoperable, and scalable, not a black box and not a bespoke artifact hidden inside a single team's workflow. That's why we are proposing a new model for structuring abstraction definitions: The Brim Abstraction Metadata Model, or BAMM.
Much like SQL gives data teams a shared grammar for querying databases, BAMM provides a shared grammar for chart abstraction. It is a formal, tool-agnostic way to describe what you want abstracted, how you want it abstracted, and how systems, whether AI or human, should interpret and combine information from clinical notes.
Why Chart Abstraction Is Broken
Abstraction underpins some of healthcare's most important work: registry submissions that drive quality improvement programs, clinical trial matching where eligibility details like ECOG scores and prior treatments live only in the notes, research protocols that depend on structured and repeatable extraction of nuanced clinical concepts, and operational workflows from readmission reviews to sepsis surveillance.
However, there is no standard way to describe the abstraction task itself.
For a deeper look at why abstraction deserves to be treated as core research infrastructure rather than a one-off project, see our earlier post: Chart Abstraction is Infrastructure, Not a Pilot.
Experienced human abstractors rely on dense manuals, institutional knowledge, months of training, and rigorous inter-rater reliability testing. This works, but it's slow, costly, and brittle. Two sites abstracting the same variable may define it differently, and when teams update a definition, the entire training process starts over.
AI introduces a different set of problems. Models require explicit instructions: not just definitions, but semantics, references, and examples. Validation demands transparency, since evaluators need to understand exactly what the system was told to do. And without a shared metadata structure, every AI tool reinvents the wheel, making it difficult to compare approaches, validate outputs, or collaborate across sites.
The result is an ecosystem where abstraction is both critically important and deeply inconsistent from institution to institution, tool to tool, team to team.
What BAMM Is
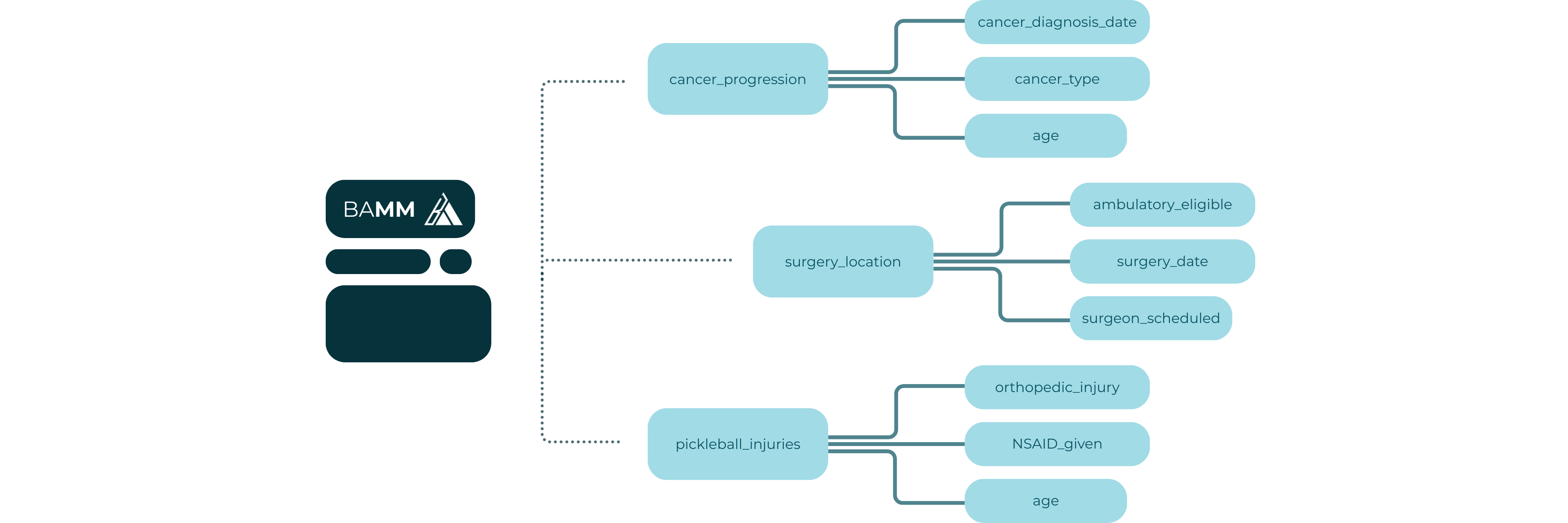
BAMM is a standardized, machine-readable schema for defining the variables you want abstracted from clinical notes. It is LLM-agnostic, tool-agnostic, and designed for both human and machine interpretation.
BAMM was built around a specific insight: the best AI abstraction systems don't try to answer high-level questions about a patient in a single pass. Instead, they abstract the smallest meaningful units first and then aggregate those granular outputs into higher-level data points. This is a map-reduce approach: the same pattern used in large-scale data processing. In the map phase, lower-scope variables run in parallel across individual notes, each one returning a granular result. In the reduce phase, higher-scope variables aggregate those results into a single patient-level value. This enables parallelism by processing hundreds of notes simultaneously rather than sequentially, improves accuracy by grounding each extraction in specific evidence rather than asking a model to synthesize a full chart at once, and makes the full pipeline auditable at every step.
BAMM formalizes that approach. Each element in the schema was chosen to address a specific failure mode in traditional abstraction, including ambiguity in definitions, inconsistency in aggregation, and opacity in execution.
The result is a framework that lets teams bring their own LLM (including small on-prem or open-source models), mix and match models across variables and projects, scale to thousands of notes and variables, and version and audit abstraction definitions over time.
What's in BAMM
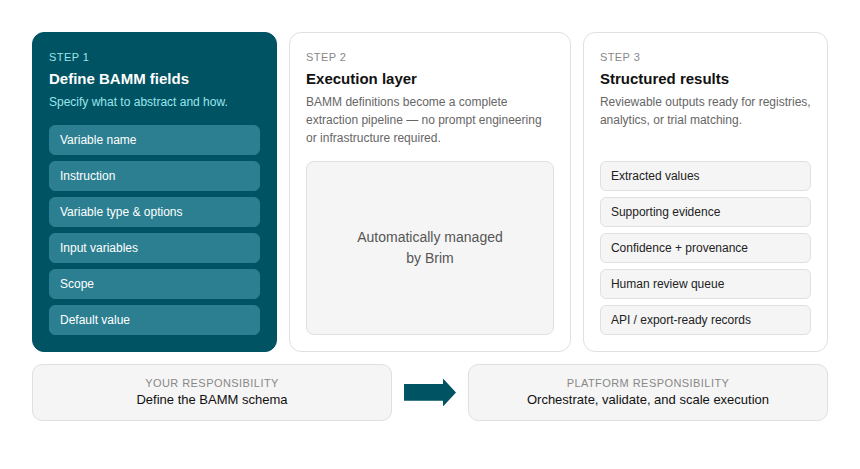
Each BAMM variable contains eight structured elements.
Variable Name. A clear, unique name for the data point being abstracted.
Instruction. A detailed description of how to extract the variable from clinical notes, including semantics, what counts as evidence, how to interpret references to other variables, and examples and edge cases. This is the core of the variable definition: what a human abstractor manual tries to capture, just made explicit and machine-readable.
Aggregation Instruction. Defines how values should be combined when synthesizing across notes or across instances of evidence. Without this, different human curators or implementation tools will aggregate differently and produce incomparable results.
Input Variables. A list of other variables that serve as prerequisites or context. This supports dependency modeling, map-reduce style execution, and integration of structured data sources alongside notes.
Variable Type. Text, integer, float, boolean, or timestamp. Ensures the returned value matches the expected type and can be validated programmatically.
Scope. Describes the level of abstraction expected. Many Labels Per Note returns all values found, One Label Per Note consolidates to one value per note, and One Label Per Patient consolidates to one value across all notes.
Options. Enumerates the accepted values for the variable's output. If empty, free text is allowed.
Default Value. The value returned when no evidence is found within the scope. This makes abstraction behavior more auditable by removing the ambiguous null.
A BAMM Variable in Practice: Cancer Registry Abstraction
To make this concrete, consider a cancer registry variable like Pathologic T Stage, representing the tumor staging value assigned based on pathology reports. Here's how BAMM might define it:
- Variable Name: Pathologic T Stage
- Instruction: Extract the pathologic T stage as documented in pathology reports. Look for explicit staging notation (e.g., pT1, pT2, pT3) in the final diagnosis or summary sections of pathology notes. Do not infer staging from tumor size alone if a stage is not explicitly assigned. If multiple pathology reports exist, prefer the most recent surgical pathology report over biopsy reports. Do not extract clinical T stage (cT). Only abstract pathologic (pT) designations.
- Aggregation Instruction: If multiple pathology reports contain a pT stage, return the value from the most recent surgical pathology report. If values conflict without a clear recency winner, return the higher stage.
- Input Variables: None
- Variable Type: Text
- Scope: One Label Per Patient
- Options: pT0, pTis, pT1, pT2, pT3, pT4, Unknown
- Default Value: Unknown
This single variable definition tells an abstraction system exactly what to look for, how to handle ambiguity, how to aggregate across multiple notes, and what to return when no evidence is found.
Orchestration: From Schema to Execution Plan
Defining a set of BAMM variables is the starting point. Turning them into a running abstraction workflow requires an orchestration layer, or a plan for how variables relate to each other and in what order they execute.
Because variables can declare dependencies on other variables through Input Variables, the full set of variables for a given abstraction task forms a directed acyclic graph. An orchestration engine reads this graph and derives an execution plan: which variables can run in parallel, which must wait for upstream results, and how outputs flow from one stage to the next.
In practice, this plays out as a pipeline. Lower-scope variables execute first and in parallel, producing granular outputs. Higher-scope variables then consume those outputs as inputs, aggregating across notes or across a patient's full history. Every intermediate result is traceable, and any variable can be re-run in isolation without restarting the full workflow.
In addition, an execution layer must handle the many complexities of working with an LLM, including:
- LLM Management: handling rate limits, token budgets, and routing across models.
- Prompt Optimization: leveraging best practices, user feedback and golden datasets to modify prompts so they continue to improve.
- Validation and QA: compare to a golden dataset, flag low-confidence values, detect hallucinations, and surface systematic errors.
- Human-in-the-loop enablement: enable humans to quickly audit values and give feedback
- Cost and performance optimization: track token usage, optimize call batching, cache results wheere appropriate, enable tradeoffs.
BAMM provides the shared vocabulary and structure that makes a good execution engine possible, but the engine itself is where the hard engineering work lives. We'll discuss this more in a future post.
Getting Started with BAMM
BAMM is designed to be adopted incrementally; you don't need to migrate an entire abstraction program at once to start benefiting from it.
The most natural entry point is a variable library. Start by documenting your existing abstraction variables in BAMM format: write out the Instruction, define the Scope, specify the Default Value. Even without an AI system behind it, this exercise surfaces inconsistencies in how your team has been defining variables and creates a shared reference that human abstractors, data scientists, and clinical stakeholders can all work from.
From there, BAMM variables can be used directly to drive LLM-based abstraction. BAMM makes it easy to adopt an abstraction tool like Brim that handles the complexities of working with an LLM at scale. It also makes it easy to try multiple tools, or use different tools in different scenarios.
BAMM also makes validation tractable. Because each variable has an explicit definition, a declared type, and a known set of options, you can build automated checks against expected output formats and compare AI results against gold-standard human abstractions systematically. Definitions that produce inconsistent outputs become visible quickly and because the definition is versioned, you can track exactly when a change was made and what effect it had.
Finally, BAMM creates the conditions for interoperability. When two institutions abstract the same variable using the same BAMM definition, their outputs are directly comparable. That opens the door to cross-site research and collaborative registry programs that would otherwise require expensive harmonization work.
Conclusion
Chart abstraction is too important to remain an ad-hoc, institution-specific process. As AI becomes deeply embedded in research, registries, and clinical workflows, we need a shared framework that ensures definitions are explicit, interoperable, and easy to validate.
BAMM is our proposal for that framework. By standardizing the metadata behind abstraction itself, we can finally scale the work that drives discovery, powers clinical operations, and improves patient outcomes across institutions, across tools, and across the AI models that will keep improving for years to come.
If your team is interested in piloting BAMM or incorporating it into a new research or quality initiative, we'd love to collaborate. Reach out at support@brimanalytics.com.